Key Concepts
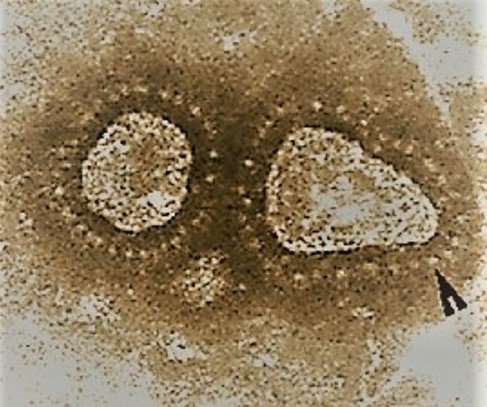
The process of identifying the presence of viruses that cause respiratory ailments. The detection of respiratory viruses greatly concerns epidemiologists and physicians. Acute respiratory diseases account for an estimated 75% of all acute morbidities in nonindustrialized countries, and most of these are caused by viruses. Upper respiratory tract infections (URTIs) are among the most common infections in children, occurring 3 to 8 times per year in children under 5 years of age, and often causing acute asthma exacerbations or acute middle ear infections. The U.S. Centers for Disease Control and Prevention's National Vital Statistics Report indicates that there are 12–32 million episodes of URTIs annually in children under 2 years of age. Viral respiratory tract infections can be caused by numerous types of viruses. These include influenza A and B, respiratory syncytial virus (RSV) types A and B, parainfluenza virus (types 1–4), numerous adenoviruses, the "common cold" viruses (including various coronaviruses and rhinoviruses), and a number of viruses discovered in the twenty-first century—metapneumovirus, various coronaviruses [including severe acute respiratory syndrome (SARS) coronavirus (Fig. 1); and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which is the virus responsible for COVID-19], bocavirus, and avian influenza virus (H5N1 and H7N9). See also: Clinical microbiology; Clinical pathology; Common cold; Coronavirus; Influenza; Novel coronavirus is declared a global pandemic; Parainfluenza virus; Respiratory syncytial virus; Respiratory system disorders; Rhinovirus; Severe acute respiratory syndrome (SARS); Virus; Virus classification
Clinical disease
All of the aforementioned virus groups can cause a full range of respiratory tract infections—from the typical mild common cold, with signs and symptoms of a runny nose and sneezing, to more severe presentations, such as pharyngitis, laryngitis, bronchitis, or pneumonia. The severity of disease varies depending on the level of immunity of the individual, and is generally more severe in immunocompromised patients [for example, those with human immunodeficiency virus (HIV); transplant recipients receiving antirejection drugs; diabetics; and the elderly]. As the immune competency of individuals and their ability to fight infection and cancer begins to decline in the fourth decade of life, the elderly patient is particularly susceptible to virus infections, which are often fatal in these circumstances. See also: Epidemiology; Immunity; Pneumonia; Public health
Traditional diagnostic methods
In clinical practice, a specific virus is often not identified due to the lack of a laboratory test that is sensitive enough to detect it. New viruses have historically accounted for a significant proportion of infections where no virus could be detected in the absence of tests. Typically, viruses have been identified by their shape and size and classified into families; for example, the coronavirus has a "crown" around its surface made up from a single protein (Fig. 1). Virology laboratories have historically diagnosed only six conventional respiratory viruses using traditional methods. These methods include, first and foremost, virus isolation in cell culture using up to four different cell lines (not all viruses grow in all cell lines). Preformed cell cultures in 15-cm-long (6-in.-long) tubes are inoculated with specimens, placed in a roller drum (where they are constantly rotated for 10 days), and viewed daily under a microscope for virus-induced cell damage, indicating the presence of a virus. In theory, culture can be sensitive enough for detection of a single living virus particle. However, at the same time, sensitivity can be lost when specific antibodies in the specimen neutralize the virus, preventing its growth (see table). See also: Cell culture
|
Methods* |
Advantages |
Disadvantages |
|---|---|---|
|
Cell culture |
Low sensitivity |
Not all viruses are culturable |
|
DFA |
Gold standard method |
Positivity rates vary by laboratory |
|
ELISA |
Provides point of care result |
Insensitive |
|
NAAT |
Highly sensitive |
Expensive |
|
M-PCR |
Detects several viruses |
Expensive |
*Abbreviations: DFA = direct fluorescent antibody; ELISA = enzyme-linked immunosorbent assay; NAAT = nucleic acid amplification test; M-PCR = multiplex polymerase chain reaction.
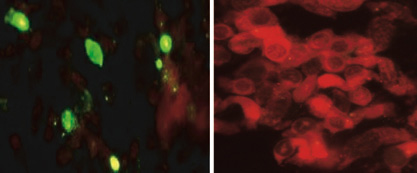
The second most important detection method has been direct fluorescent antibody (DFA) staining for the presence of virus-infected cells. This method involves collecting epithelial cells from a nasopharyngeal swab, fixing them to a glass microscope slide, staining with individual antibodies labeled with a fluorescent tag, and viewing the slide with a fluorescent microscope (Fig. 2). This method has sensitivities ranging from 65% to 90% for the six viruses commonly detected. Sensitivity can be compromised if the specimen is collected too late in the course of infection when the number of cells containing viral proteins is diminished. The third most commonly used method is shell vial culture. This involves inoculating an aliquot of the specimen onto a preformed cell monolayer in a small vial containing a mixture of two susceptible cells, which is then centrifuged to enhance virus attachment and entry. The centrifugation-assisted inoculation of the cells increases the amount of viral proteins produced, allowing staining to be performed at 24–48 h and thus providing a test result to be obtained significantly earlier than the 7–10 days necessary for traditional cell culture. Rapid enzyme-linked immunosorbent assays (ELISAs), in which a monoclonal antibody conjugated to an enzyme is used to rapidly detect and quantify the presence of an antigen in a sample, have been developed as bedside tests for influenza and RSV.
Molecular methods
The aforementioned traditional methods have been the cornerstone for diagnosis used by virology laboratories around the world. The introduction of nucleic acid amplification tests (NAATs) for respiratory viruses starting in the late 1980s heralded a new era in diagnosing respiratory virus infections. The first NAAT for respiratory viruses was developed for influenza and used a nucleic acid amplification method called polymerase chain reaction (PCR), developed in 1983 by Kary B. Mullis, who was later awarded the Nobel Prize in Chemistry in 1993. Within a decade, NAATs were developed for all of the respiratory viruses, and most used PCR; however, other amplification schemes, such as nucleic acid–sequence-based amplification (NASBA), strand displacement amplification (SDA), transcription-mediated amplification (TMA), and loop-mediated isothermal amplification (LAMP), have also been used. For all NAATs, the total nucleic acid is first extracted from the respiratory tract specimen using a variety of methods, and the viral ribonucleic acid (RNA) is copied into a complementary deoxyribonucleic acid (cDNA) using an enzyme called reverse transcriptase. The cDNA is then amplified by PCR using virus-specific oligonucleotide primers, resulting in a billion copies of DNA that can be easily detected by a variety of common laboratory methods. Following the emergence of new human respiratory viruses in the twenty-first century, there has been a need for new diagnostic tests to detect these viral pathogens, and NAAT filled this need. Early comparisons of molecular and traditional methods clearly indicated that the molecular methods were more sensitive than the traditional methods, often diagnosing up to 30% additional infected patients. Molecular testing methods also provided test results for clinicians often within 1 day (as compared with 2–5 days needed for traditional methods), thus improving their management of patients. See also: Nucleic acid; Oligonucleotide; Polymerase chain reaction (PCR)
The next major advance in diagnostics was the development of multiplex PCR (M-PCR) for the detection of several different viruses in a single test. M-PCR uses multiple oligonucleotide primers, with one pair for each virus to be detected. Because M-PCR will detect several different viruses, a method is required to identify which virus is present in the specimen. This is done using a microarray (a collection of several different DNA oligonucleotides) that is either spotted onto microscope slides or cartridges (gene chips) or immobilized onto microspheres (microfluidic arrays) that are each uniquely labeled using a mixture of fluorescent dyes and identified by lasers. Each element in the array (a spot or microsphere) consists of a unique oligonucleotide (representing a unique virus) that will bind individual PCR products for each virus type or subtype. A positive specimen will generate an amplification product that is hybridized to one of the elements of the microarray and detected by a laser. One M-PCR called the xTAG respiratory viral panel (RVP) is used as an in vitro diagnostic device. This test was designed to identify 20 different respiratory virus types and subtypes, and it uses more than 30 primers for target amplification and identification. Although M-PCR tests are slightly more expensive than single NAATs, they have the advantage of being able to detect many different viruses in a single test, as well as being able to detect dual infections occurring in about 10% of patients and even triple infections that are not often seen with traditional methods. NAATs using M-PCR and microarray technologies offer unprecedented power for the laboratory, and are well on the way to becoming the pillars of diagnostic virology for the present century.





